import mikeio
ds = mikeio.read("../data/gebco_sound.dfs2")
ds<mikeio.Dataset>
dims: (time:1, y:264, x:216)
time: 2020-05-15 11:04:52 (time-invariant)
geometry: Grid2D (ny=264, nx=216)
items:
0: Elevation <Total Water Depth> (meter)A dfs2 file is also called a grid series file. Values in a dfs2 file are ‘element based’, i.e. values are defined in the centre of each grid cell.
<mikeio.Dataset>
dims: (time:1, y:264, x:216)
time: 2020-05-15 11:04:52 (time-invariant)
geometry: Grid2D (ny=264, nx=216)
items:
0: Elevation <Total Water Depth> (meter)The most convenient way to subset in space is to use the sel method, which returns a new (smaller) dataset, which can be further processed or written to disk using the to_dfs method.
<mikeio.Grid2D>
x: [12.2, 12.21, ..., 13.1] (nx=216, dx=0.004167)
y: [55.2, 55.21, ..., 56.3] (ny=264, dy=0.004167)
projection: LONG/LAT<mikeio.Grid2D>
x: [12.5, 12.5, ..., 12.99] (nx=120, dx=0.004167)
y: [55.5, 55.5, ..., 55.99] (ny=120, dy=0.004167)
projection: LONG/LATIn order to specify an open-ended subset (i.e. where the end of the subset is the end of the domain), use None as the end of the slice.
The spatial information is available in the geometry attribute (accessible from Dfs2, Dataset, and DataArray), which in the case of a dfs2 file is a Grid2D geometry.
<mikeio.Grid2D>
x: [12.2, 12.21, ..., 13.1] (nx=216, dx=0.004167)
y: [55.2, 55.21, ..., 56.3] (ny=264, dy=0.004167)
projection: LONG/LATGrid2D’s primary properties and methods are:
xnxdxynydyoriginprojectionxybboxcontains()find_index()isel()to_mesh()See API specification for details.
Use interp() to extract values at a specific coordinate:
To create a dfs2 file from scratch:
Grid2D geometryDataArrayData arrays must have shape (n_timesteps, ny, nx) for time-varying data or (ny, nx) for static data. The y-axis must increase south-to-north and the x-axis west-to-east, matching the CF conventions.
For complete examples of converting external data to dfs2, see Dfs2-Bathymetry (NetCDF → static dfs2) and Dfs2-GFS (NetCDF → multi-item time-varying dfs2).
<mikeio.Grid2D>
x: [12, 13, 14] (nx=3, dx=1)
y: [55, 56, ..., 58] (ny=4, dy=1)
projection: LONG/LAT<mikeio.DataArray>
name: Elevation
dims: (time:2, y:4, x:3)
time: 2020-01-01 00:00:00 - 2020-01-02 00:00:00 (2 records)
geometry: Grid2D (ny=4, nx=3)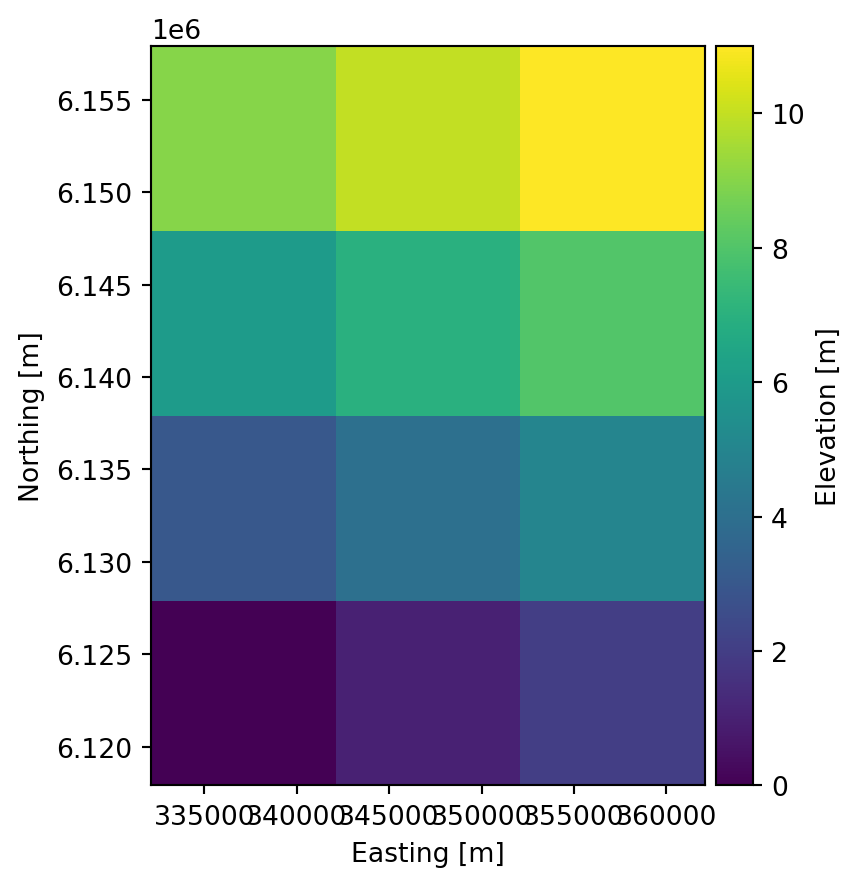
Cleanup: