from modelskill.network import Network
network = Network.from_res1d(path_to_res1d)
network<Network>
Reaches: 118
Nodes: 259
Quantities: ['WaterLevel', 'Discharge']
Time: 1994-08-07 16:35:00 - 1994-08-07 18:35:00A Network represents a 1D pipe or river network as a directed graph: nodes hold timeseries data (e.g. water level at a junction) and reaches carry the topology and reach length between them. Break points along a reach (e.g. cross-section chainages) are supported as observation locations too.
The typical workflow is:
Network → NetworkModelResult → match() → ComparerYou can build a Network object by loading it from a supported network format.
Currently, the only supported format is mikeio.Res1D.
The quickest way to get a Network is from the path to a MIKE 1D result file:
<Network>
Reaches: 118
Nodes: 259
Quantities: ['WaterLevel', 'Discharge']
Time: 1994-08-07 16:35:00 - 1994-08-07 18:35:00or a mikeio1d.Res1D that has already been opened:
A Res1D network contains multiple levels that are unified into a generic network structure as depicted in the image below. The image introduces concepts like find, recall and boundary which are explained in the following sections.
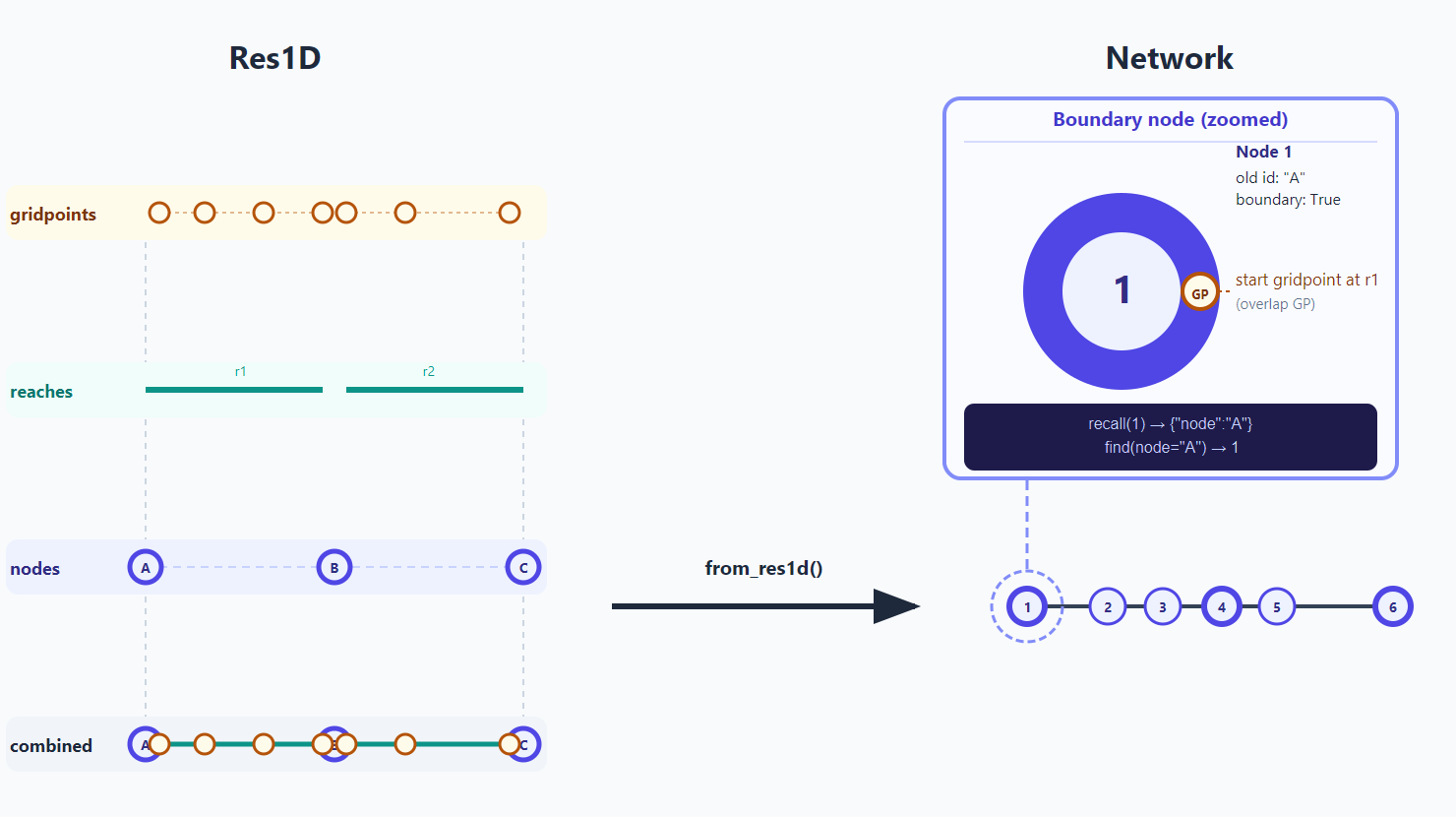
find()/recall() round-trip lookups.Large Res1D files can contain thousands of nodes and gridpoints. Loading all of that data into memory is slow and may cause memory issues — especially when you only need the timeseries at a handful of nodes where observations exist.
from_res1d accepts two optional arguments to restrict what gets loaded:
| Argument | Type | Effect |
|---|---|---|
nodes |
None | str | list[str] |
Control which nodes have timeseries data loaded. None (default) loads all nodes; [] skips all node data; a name or list loads only those nodes. |
reaches |
None | str | list[str] |
Control which reaches have intermediate gridpoint data populated. None (default) loads everything; [] skips all gridpoints; a name or list of names loads only those reaches. |
Selective loading only controls which timeseries are held in memory. The full network topology (nodes, reaches, lengths) is always constructed so that find(), recall(), and graph algorithms still work on the complete network.
The most memory-efficient setup — useful when you only care about specific junction nodes — is to pass the node IDs you need and skip all intermediate gridpoints with reaches=[]:
<Network>
Reaches: 118
Nodes: 259
Quantities: ['WaterLevel']
Time: 1994-08-07 16:35:00 - 1994-08-07 18:35:00If you also need gridpoint data along a particular reach, pass its name (or a list of names):
<Network>
Reaches: 118
Nodes: 259
Quantities: ['WaterLevel', 'Discharge']
Time: 1994-08-07 16:35:00 - 1994-08-07 18:35:00When only some nodes are loaded, to_dataframe() and to_dataset() only contain columns for those nodes — the rest are graph-connected but data-free:
| node | 134 | 203 |
|---|---|---|
| time | ||
| 1994-08-07 16:35:00 | 194.074997 | 194.852997 |
| 1994-08-07 16:36:01.870000 | 194.074997 | 194.853577 |
| 1994-08-07 16:37:07.560000 | 194.074997 | 194.853760 |
| 1994-08-07 16:38:55.828000 | 194.074997 | 194.853836 |
| 1994-08-07 16:39:55.828000 | 194.074997 | 194.853867 |
The network exposes a networkx.Graph so you can use any NetworkX algorithm or plotting function directly:
| node | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | ... | 249 | 250 | 251 | 252 | 253 | 254 | 255 | 256 | 257 | 258 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| quantity | WaterLevel | WaterLevel | Discharge | WaterLevel | Discharge | WaterLevel | WaterLevel | Discharge | WaterLevel | WaterLevel | ... | Discharge | WaterLevel | Discharge | WaterLevel | Discharge | Discharge | Discharge | WaterLevel | Discharge | Discharge |
| time | |||||||||||||||||||||
| 1994-08-07 16:35:00.000 | 195.441498 | 194.661499 | 0.000006 | 195.931503 | 0.000004 | 193.550003 | 193.550003 | 0.000000 | 195.801498 | 195.703003 | ... | 0.000005 | 194.511505 | 0.000013 | 194.581497 | 0.000003 | 0.000002 | 0.000031 | 193.779999 | 0.0 | 0.0 |
| 1994-08-07 16:36:01.870 | 195.441605 | 194.661621 | 0.000006 | 195.931595 | 0.000004 | 193.550140 | 193.550064 | 0.000008 | 195.801498 | 195.703171 | ... | 0.000005 | 194.511841 | 0.000010 | 194.581497 | 0.000003 | 0.000002 | 0.000031 | 188.479996 | 0.0 | 0.0 |
| 1994-08-07 16:37:07.560 | 195.441620 | 194.661728 | 0.000006 | 195.931625 | 0.000004 | 193.550232 | 193.550156 | 0.000016 | 195.801498 | 195.703400 | ... | 0.000005 | 194.511795 | 0.000010 | 194.581497 | 0.000003 | 0.000002 | 0.000033 | 188.479996 | 0.0 | 0.0 |
| 1994-08-07 16:38:55.828 | 195.441605 | 194.661926 | 0.000006 | 195.931656 | 0.000004 | 193.550369 | 193.550308 | 0.000022 | 195.801498 | 195.703690 | ... | 0.000005 | 194.511581 | 0.000009 | 194.581497 | 0.000003 | 0.000002 | 0.000037 | 188.479996 | 0.0 | 0.0 |
| 1994-08-07 16:39:55.828 | 195.441605 | 194.661972 | 0.000006 | 195.931656 | 0.000004 | 193.550430 | 193.550369 | 0.000024 | 195.801498 | 195.703827 | ... | 0.000005 | 194.511505 | 0.000009 | 194.581497 | 0.000003 | 0.000002 | 0.000039 | 188.479996 | 0.0 | 0.0 |
5 rows × 259 columns
| node | 0 | 1 | 3 | 5 | 6 | 8 | 9 | 11 | 12 | 14 | ... | 235 | 237 | 239 | 241 | 244 | 246 | 248 | 250 | 252 | 256 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| time | |||||||||||||||||||||
| 1994-08-07 16:35:00.000 | 195.441498 | 194.661499 | 195.931503 | 193.550003 | 193.550003 | 195.801498 | 195.703003 | 197.072006 | 196.962006 | 197.351501 | ... | 196.272995 | 196.113007 | 196.322006 | 196.401993 | 196.851501 | 196.891495 | 196.601501 | 194.511505 | 194.581497 | 193.779999 |
| 1994-08-07 16:36:01.870 | 195.441605 | 194.661621 | 195.931595 | 193.550140 | 193.550064 | 195.801498 | 195.703171 | 197.072006 | 196.962051 | 197.351501 | ... | 196.272995 | 196.113007 | 196.322037 | 196.402023 | 196.851501 | 196.891495 | 196.601501 | 194.511841 | 194.581497 | 188.479996 |
| 1994-08-07 16:37:07.560 | 195.441620 | 194.661728 | 195.931625 | 193.550232 | 193.550156 | 195.801498 | 195.703400 | 197.072006 | 196.962082 | 197.351501 | ... | 196.272995 | 196.113007 | 196.322052 | 196.402039 | 196.851501 | 196.891495 | 196.601501 | 194.511795 | 194.581497 | 188.479996 |
| 1994-08-07 16:38:55.828 | 195.441605 | 194.661926 | 195.931656 | 193.550369 | 193.550308 | 195.801498 | 195.703690 | 197.072006 | 196.962112 | 197.351501 | ... | 196.272995 | 196.113007 | 196.322067 | 196.402069 | 196.851501 | 196.891495 | 196.601501 | 194.511581 | 194.581497 | 188.479996 |
| 1994-08-07 16:39:55.828 | 195.441605 | 194.661972 | 195.931656 | 193.550430 | 193.550369 | 195.801498 | 195.703827 | 197.072006 | 196.962128 | 197.351501 | ... | 196.272995 | 196.113007 | 196.322067 | 196.402069 | 196.851501 | 196.891495 | 196.601501 | 194.511505 | 194.581497 | 188.479996 |
5 rows × 130 columns
After construction, nodes are re-labelled as integers. Use find() to go from original coordinates to the integer ID and recall() to go back.
When creating NodeObservation objects for skill assessment you generally do not need to call find(). You can pass the original string ID as node=, and NetworkModelResult will resolve it for you during matching. For breakpoints, use at=(reach, distance) rather than node=. See Skill assessment workflow for details.
Node '117' → integer id 51
{'node': '117'}Break point (94l1, 21.285) → integer id 245
{'reach': '94l1', 'distance': 21.2852238539205}NodeObservation accepts a file path directly; the observation name is taken from the filename.
The at= argument can be specified in three ways, depending on what information you have at hand.
In MIKE 1D, “nodes” are the named connection points in the 1D network (manholes, junctions, outfalls) while “reaches” are the pipe or channel segments that connect them. The modelskill API uses NodeObservation more broadly: the at= argument accepts either a node ID (int or string) or a (reach_id, distance) breakpoint tuple that identifies a specific chainage along a reach. If you only need a reach-level quantity (uniform across the reach), use ReachObservation with reach= instead.
Pass the original node identifier from the source format (e.g. the Res1D node name) as a plain string. The NetworkModelResult resolves it to the correct integer ID at match time, so you do not need to call network.find() yourself:
| n | bias | rmse | urmse | mae | cc | si | r2 | |
|---|---|---|---|---|---|---|---|---|
| observation | ||||||||
| network_sensor_1 | 109 | 0.700685 | 0.810246 | 0.406865 | 0.724734 | 0.726882 | 0.002095 | -2.245889 |
| network_sensor_2 | 80 | 0.430548 | 0.462543 | 0.169040 | 0.431153 | 0.550352 | 0.000873 | -4.548691 |
Resolution happens inside ms.match(). If the string is not found in the network’s alias map a ValueError is raised with a clear message indicating which alias could not be resolved.
(reach, distance) tupleWhen your observation sits at a chainage along a reach rather than at a named junction node, you can use the at argument and pass a (reach_id, distance) tuple. The NetworkModelResult looks up the corresponding breakpoint at match time:
The tuple form is equivalent to calling network.find(reach="94l1", distance=21.285) beforehand and is resolved during matching.
Breakpoint distances are matched with a tolerance of 1 × 10⁻³ (i.e. ±0.001 in whatever distance units the network uses). This means that small floating-point discrepancies between the distance you type and the value stored in the network are handled gracefully. If no breakpoint falls within that tolerance a ValueError is raised.
Some physical quantities — such as discharge in a pipe — are conceptually uniform across the whole reach, even though the model stores values at individual breakpoints along the reach. ReachObservation lets you associate a timeseries with a named reach without having to identify a specific breakpoint.
When matched against a NetworkModelResult, modelskill automatically extracts model data from an arbitrary breakpoint that belongs to the given reach and verifies that all breakpoints on the reach carry equivalent values.
<ReachObservation>: Q_94l1
Time: 1994-08-07 16:35:06.721400 - 1994-08-07 18:35:01.353900
Quantity: []Pass the observation to ms.match() exactly as you would a NodeObservation. modelskill resolves which breakpoint to use automatically:
| n | bias | rmse | urmse | mae | cc | si | r2 | |
|---|---|---|---|---|---|---|---|---|
| observation | ||||||||
| Q_94l1 | 109 | -194.191817 | 194.192311 | 0.438201 | 194.191817 | 0.563474 | 0.002256 | -186449.466671 |
Use ReachObservation when your measured quantity is representative of the whole reach (e.g. discharge, which is constant along a reach in steady flow). If you need to compare a quantity that varies spatially along the reach (e.g. water level at a specific chainage), use a NodeObservation with a (reach, distance) tuple instead (see Option B above).
In case you have your network data in a format that is not included in Building a Network, you can assemble a Network object by subclassing the abstract base classes NetworkNode and NetworkReach.
NetworkNode requires three properties: id, data, and boundary. NetworkReach requires five: id, start, end, length, and breakpoints.
The following is a simple implementation example:
import pandas as pd
import numpy as np
from typing import Any
from modelskill.network import NetworkNode, NetworkReach, Network
class ExampleNode(NetworkNode):
"""Node backed by an in-memory DataFrame, e.g. model output."""
def __init__(self, node_id: str, data: pd.DataFrame):
self._id = node_id
self._data = data
@property
def id(self) -> str:
return self._id
@property
def data(self) -> pd.DataFrame:
return self._data
@property
def boundary(self) -> dict[str, Any]:
return {}
class ExampleReach(NetworkReach):
"""Reach connecting two nodes with a given length."""
def __init__(
self, reach_id: str, start: NetworkNode, end: NetworkNode, length: float,
breakpoints: list | None = None,
):
self._id = reach_id
self._start = start
self._end = end
self._length = length
self._breakpoints = breakpoints or []
@property
def id(self) -> str:
return self._id
@property
def start(self) -> NetworkNode:
return self._start
@property
def end(self) -> NetworkNode:
return self._end
@property
def length(self) -> float:
return self._length
@property
def breakpoints(self) -> list:
return self._breakpointsThe three abstract properties that every NetworkNode subclass must implement are id, data and boundary. If boundary is not relevant for your use case, define the property to return an empty dictionary, as in the example above. Similarly, a NetworkReach with no intermediate points can return an empty breakpoints list.
from modelskill.network import Network
# df1, df2 and df3 are DataFrame objects that are loaded in memory
node_s1 = ExampleNode("sensor_1", df1)
node_s2 = ExampleNode("sensor_2", df2)
node_s3 = ExampleNode("sensor_3", df3)
reach1 = ExampleReach("r1", node_s1, node_s2, length=500.0)
reach2 = ExampleReach("r2", node_s2, node_s3, length=300.0)
network = Network(reaches=[reach1, reach2])
network<Network>
Reaches: 2
Nodes: 3
Quantities: ['WaterLevel']
Time: 1994-08-07 16:00:00 - 1994-08-07 18:59:00Break points represent intermediate chainage locations on a reach (e.g. cross-sections). Subclass ReachBreakPoint the same way — implement id (a (reach_id, distance) tuple) and data:
from modelskill.network import ReachBreakPoint
class ExampleBreakPoint(ReachBreakPoint):
def __init__(self, reach_id: str, distance: float, data: pd.DataFrame):
self._id = (reach_id, distance)
self._data = data
@property
def id(self):
return self._id
@property
def data(self):
return self._data
# df4 is a DataFrame object that has been loaded in memory
bp = ExampleBreakPoint("r1", 200.0, df4)
reach1 = ExampleReach("r1", node_s1, node_s2, length=500.0, breakpoints=[bp])
reach2 = ExampleReach("r2", node_s2, node_s3, length=300.0)
network = Network(reaches=[reach1, reach2])